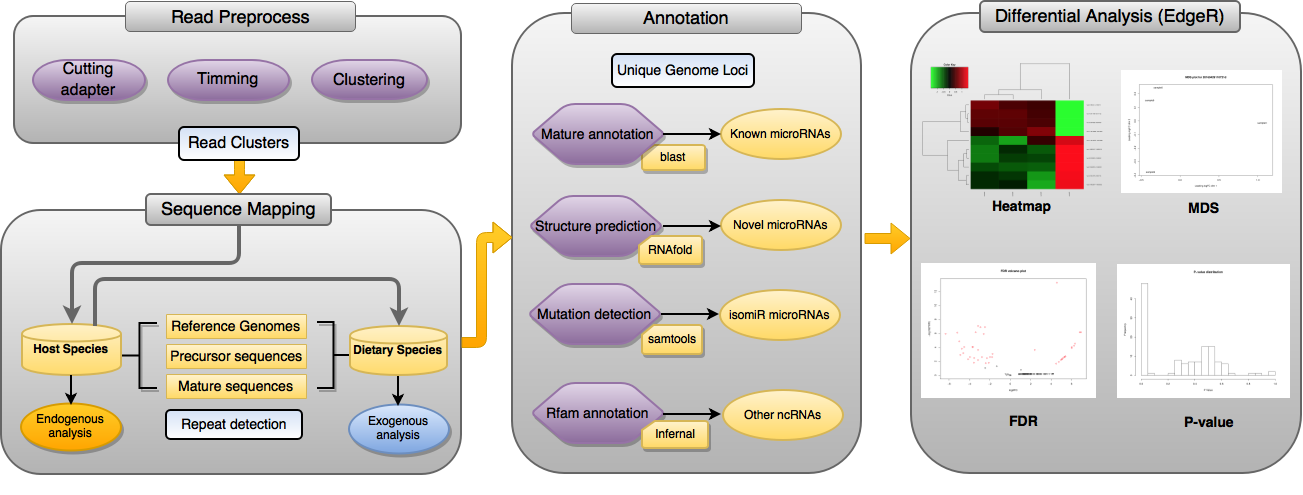
Small RNA sequencing as the most widely used tool for microRNA discovery shows great potential to study microRNA cross-species transport in an efficient manner. With the increased appreciation on dietary microRNA and its far-reaching implication in human health, research interests in exogenous microRNAs regarding their bioavailability, mechanisms of cross-species transport, and roles in major cellular biological processes has increasingly soared. Here we present miRDis, a new pipeline for both endogenous and exogenous microRNAs analysis based on deep sequencing data. Specifically, we deployed a web service that supports the annotation and expression profiling of known host microRNAs and the discovery of novel microRNAs and exogenous microRNAs resulted from dietary uptake. As a case study, we analyzed in-house sequencing data on human plasma with milk feeding. 227 human microRNAs were detected in the plasma samples where 44 microRNAs show elevated expression after milk intake. By examining the bovine specific sequences, evidence strongly indicates that three bovine microRNAs (bta-miR-378, -181*, and -150) are present in human plasma due to the dietary uptake. Further evaluation using public sequencing data demonstrated that miRDis outperforms state-of-the-art tools in both microRNA detection and quantification. The miRDis web server is available at: http://sbbi.unl.edu/miRDis/index.php.