1. General introduction
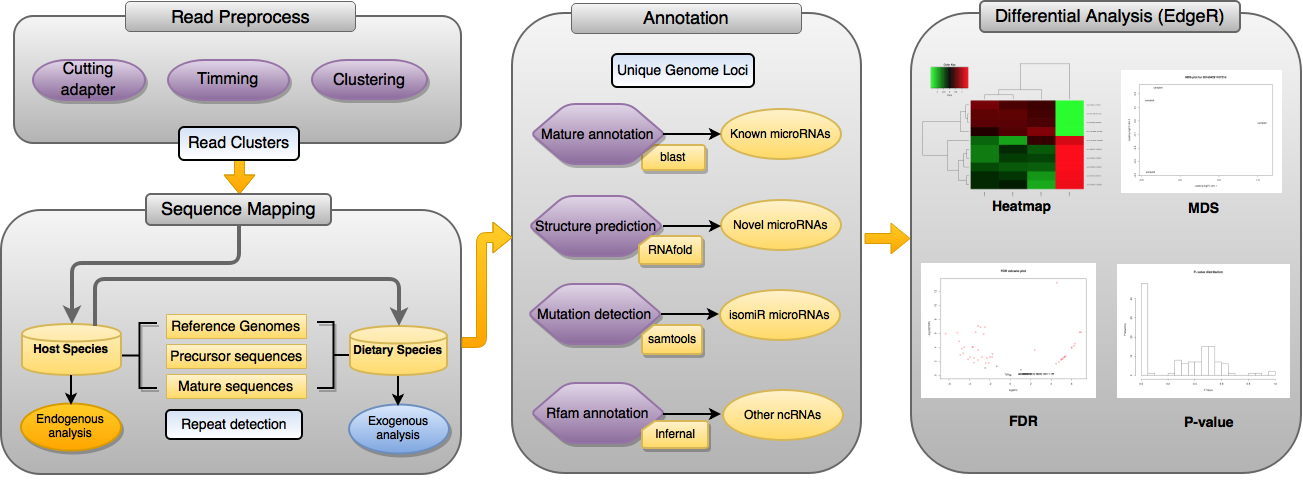
In this work, we present miRDis, a web-based small RNA sequencing analytical pipeline that display the following key features (i) systemically annotation of known miRNAs and other non-coding RNAs based on read mapped regions (ii) prediction of novel miRNAs and non-coding RNAs through assigning ambiguous mapped reads to unique genome region with well predicted RNA structure (iii) detection of candidate exogenous miRNA from dietary related species (v) support of the comparative analysis through integration of the differential analysis package. Through a simple graphical interface, users can use the full analysis of this one-stop tool for miRNA sequencing data analysis through minimal parameter settings. The tabular and graphical output contains detailed reports on the read alignment, annotation, and statistics.